from pathlib import Path
from skimage.io import imread
from skimage.draw import ellipse
from skimage.exposure import rescale_intensity
from skimage.filters import gaussian
from skimage.filters import threshold_otsu
from skimage.measure import find_contours
from skimage.measure import label
from skimage.measure import regionprops
from skimage.measure import regionprops_table
from skimage.transform import rotate
import warnings
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd31 Porosity quantification
In this notebook we implement a simple segmentation routine for porosity quantification.
Workflow is mostly based on scikit-image package.
You should run it sequentially and check quality of outputs at each step.
31.1 Required tools
media = Path(".").resolve().parent / "media" / "samples-porosity"
media.exists()Truedef display_img(img, cmap="gray", no_ticks=True, **kw):
""" Display an image in a standard way with no borders. """
fig, ax = plt.subplots()
ax.imshow(img, cmap=cmap, **kw)
if no_ticks:
ax.axis("off")
plt.subplots_adjust(left=0, right=1, top=1, bottom=0)31.2 Load and inspect initial image
Consider providing the relative or full path to target image.
# fname = media / "500004.JPG"
# fname = media / "700005.JPG"
fname = media / "800009.JPG"
fname.exists()TrueImage is loaded in gray-scale, i.e. as a 2-D array, for later thresholding.
img0 = imread(fname, as_gray=True)
display_img(img0)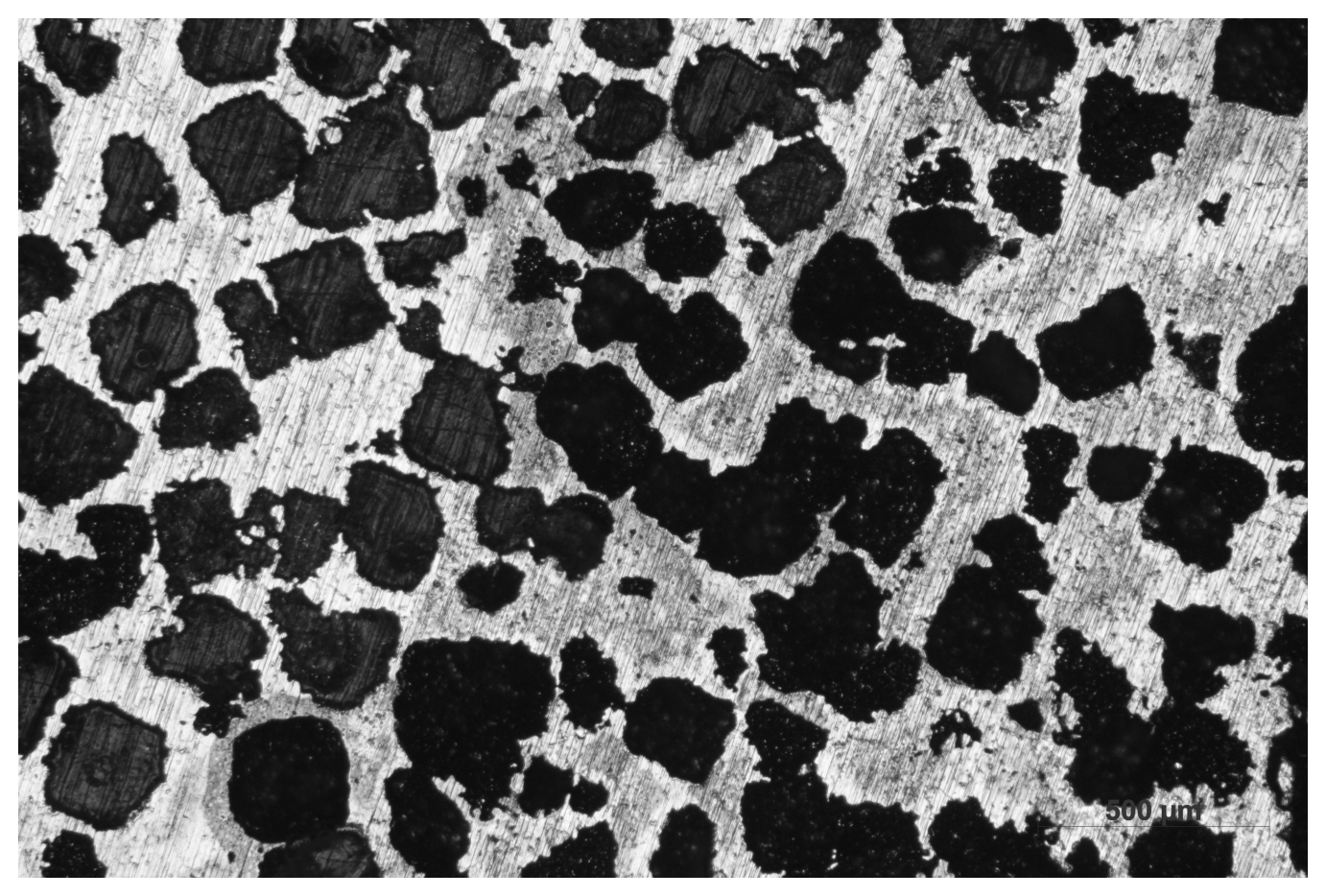
31.3 Blurring
Because of grinding and etching defects, it is important to apply an initial blur to the image.
You can control this by changing the value of sigma, the variance of the filter.
NOTE: the sigma value you choose here will impact the value of threshold img_lo you will use later. If Otsu automated thresholding works properly, you do not need to tweak this parameter and the associated img_lo. This is provided for cases where Otsu fails to automatically get pore boundaries.
img1 = gaussian(img0, sigma=15)
display_img(img1)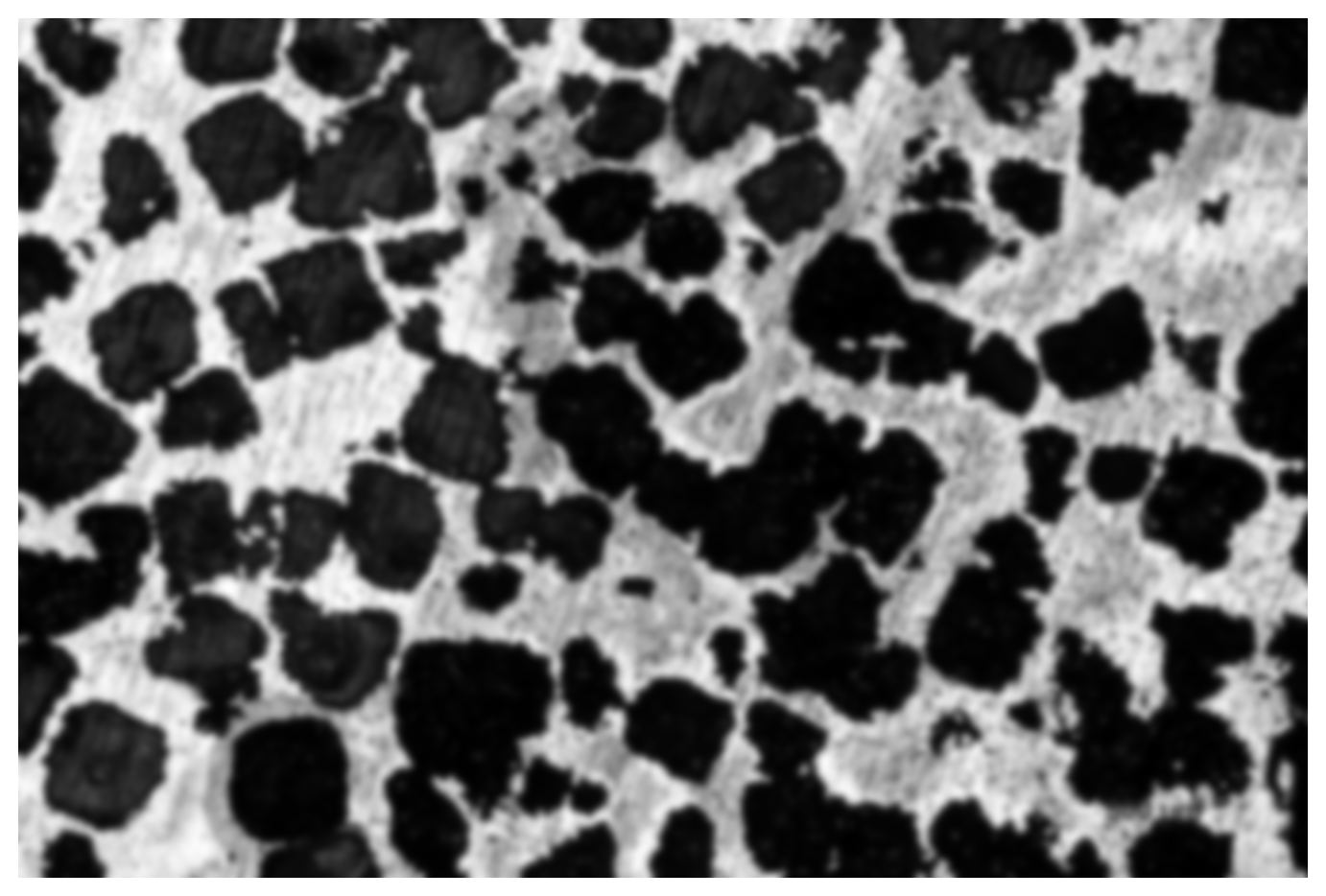
31.4 Rescaling
Rescaling with standardize the range of image for later binary selection.
There is no need to tweak the next cell.
img2 = rescale_intensity(img1, out_range=(-1, 1))
display_img(img2)
31.5 Manual thresholding
Using a cut-off lower threshold we can capture the pores.
You need to manually edit parameter img_lo, which should be close to -1 because image intensity has been scaled to \(I\in[-1,1]\).
img_lo = -0.4
img3 = np.ones_like(img2)
img3[img2 < img_lo] = 0
display_img(img3)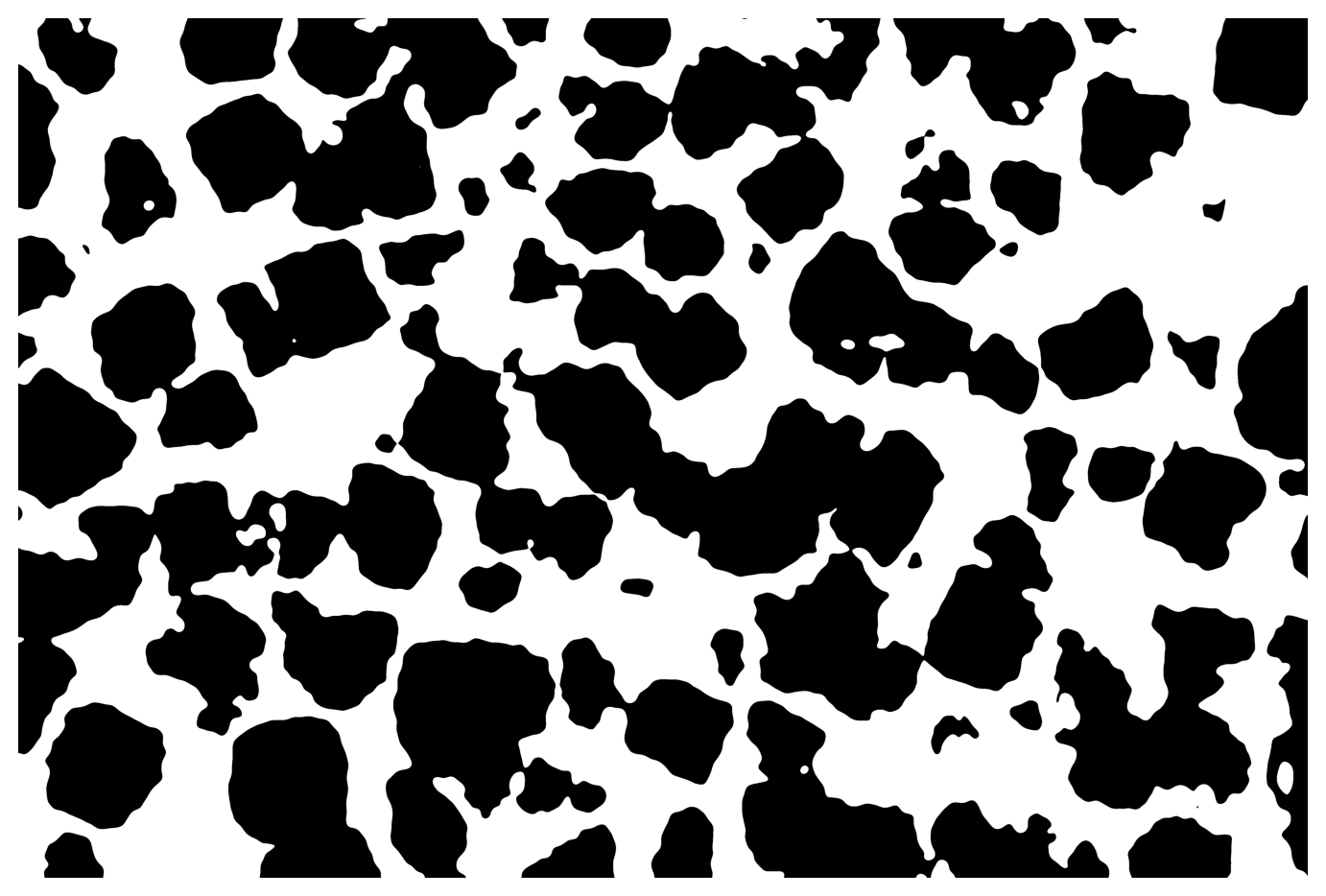
31.6 Automatic thresholding
The following is based on this example provided by scikit-image.
img4 = img1 > threshold_otsu(img1)
display_img(img4)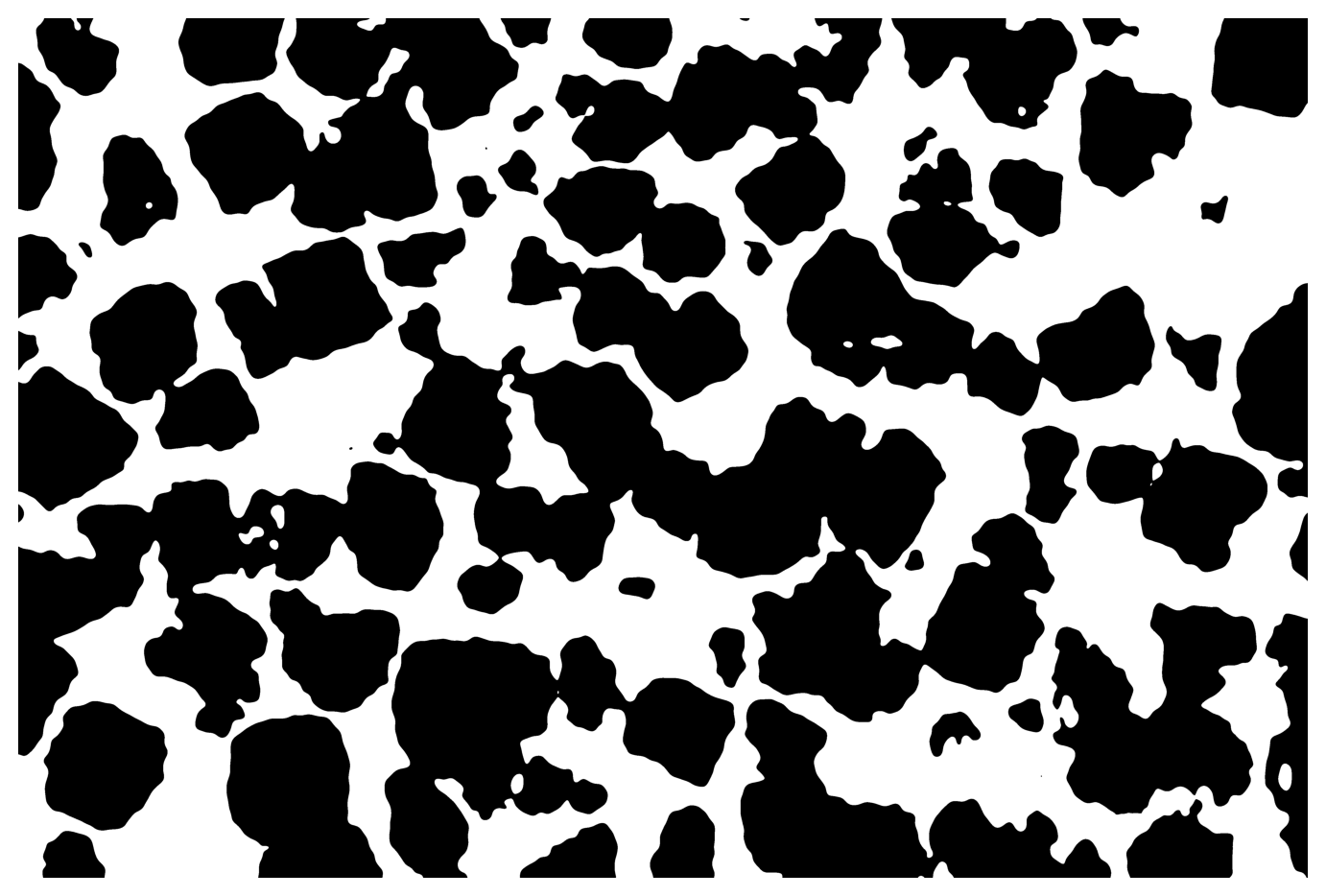
31.7 Validation and results
We find the countours of pores for display with the original image.
It is recommended you check the aspect to confirm the quality of results.
porosity_manu = 100 * (1 - img3.sum() / img0.size)
porosity_auto = 100 * (1 - img4.sum() / img0.size)
cmanu = find_contours(img3, 0.99)
cauto = find_contours(img4, 0.99)
fig, ax = plt.subplots(1, 2, figsize=(12, 6))
ax[0].set_title(f"Manual porosity of {porosity_manu:.1f}%")
ax[1].set_title(f"Automated porosity of {porosity_auto:.1f}%")
ax[0].imshow(img0, cmap="gray")
ax[1].imshow(img0, cmap="gray")
for c in cmanu:
ax[0].plot(c[:, 1], c[:, 0], color="r", linewidth=0.5)
for c in cauto:
ax[1].plot(c[:, 1], c[:, 0], color="r", linewidth=0.5)
ax[0].axis("off")
ax[1].axis("off")
fig.tight_layout()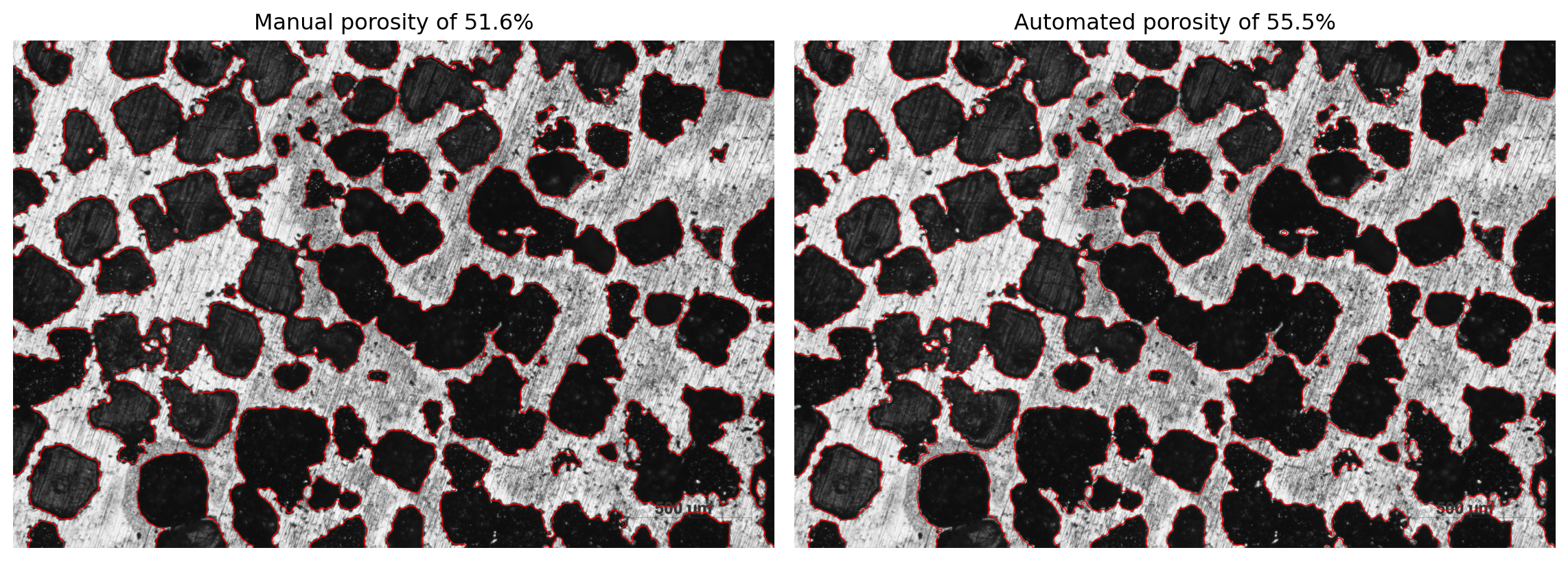
31.8 Workflow wrap-up
def porosity_workflow(fname, sigma=15):
""" Mostly automated quantification (play with sigma). """
img0 = imread(fname, as_gray=True)
img1 = gaussian(img0, sigma=sigma)
img2 = rescale_intensity(img1, out_range=(-1, 1))
img4 = img1 > threshold_otsu(img1)
porosity = 100 * (1 - img4.sum() / img0.size)
contours = find_contours(img4, 0.99)
plt.close("all")
fig, ax = plt.subplots(1, 1, figsize=(6, 5))
ax.set_title(f"Automated porosity of {porosity:.1f}%")
ax.imshow(img0, cmap="gray")
for c in contours:
ax.plot(c[:, 1], c[:, 0], color="r", linewidth=1)
ax.axis("off")
fig.tight_layout()porosity_workflow(fname)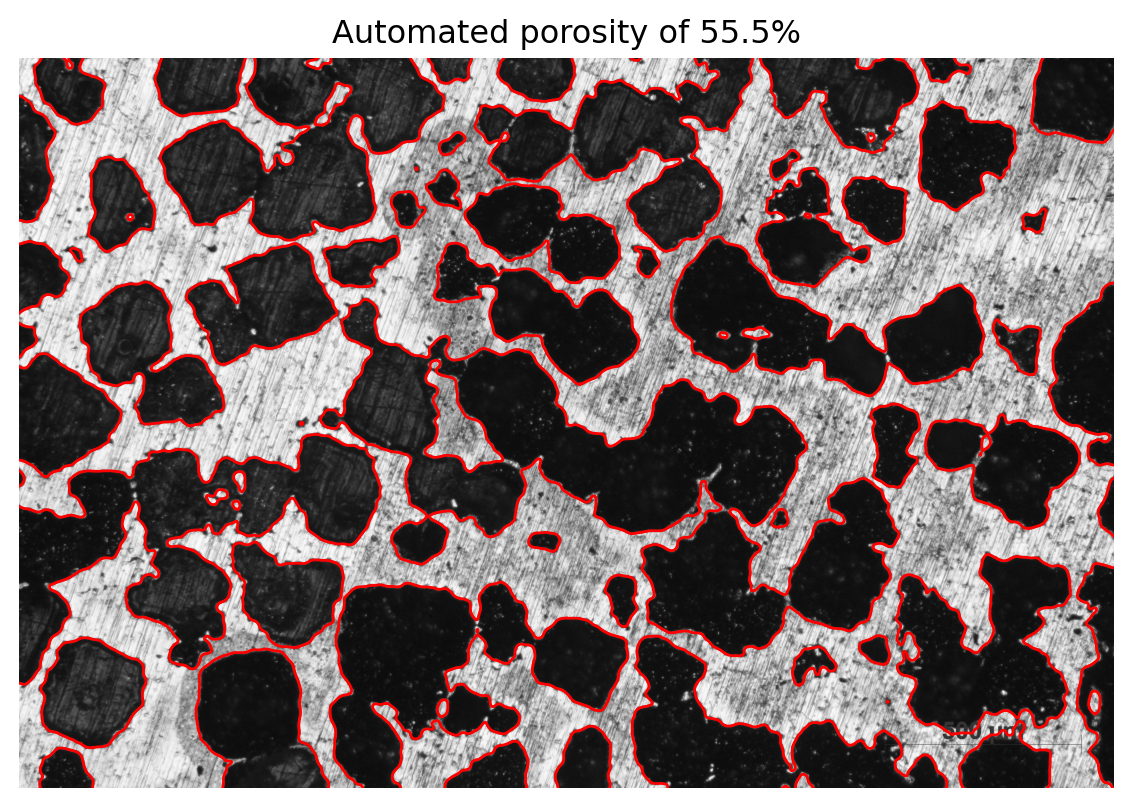
31.9 Region properties bonus
def plot_region(ax, props, cutoff):
""" Display a region with borders and equivalent ellipse. """
if props.area < cutoff:
return
y0, x0 = props.centroid
theta = props.orientation
x1 = x0 + np.cos(theta) * props.axis_minor_length / 2
y1 = y0 - np.sin(theta) * props.axis_minor_length / 2
x2 = x0 - np.sin(theta) * props.axis_major_length / 2
y2 = y0 - np.cos(theta) * props.axis_major_length / 2
# TODO test if not at border
ax.plot((x0, x1), (y0, y1), "-r", linewidth=1)
ax.plot((x0, x2), (y0, y2), "-r", linewidth=1)
ax.plot(x0, y0, ".g", markersize=15)
minr, minc, maxr, maxc = props.bbox
bx = (minc, maxc, maxc, minc, minc)
by = (minr, minr, maxr, maxr, minr)
ax.plot(bx, by, "-b", linewidth=1)def plot_all_regions(img, contours, regions, cutoff):
""" Plot all detected regions in an image. """
plt.close("all")
fig, ax = plt.subplots(1, 1, figsize=(6, 5))
# ax.set_title(f"Automated porosity of {porosity:.1f}%")
ax.imshow(img, cmap="gray")
for c in contours:
ax.plot(c[:, 1], c[:, 0], color="y", linewidth=1)
for props in regions:
plot_region(ax, props, cutoff)
ax.axis("off")
plt.subplots_adjust(left=0, right=1, top=1, bottom=0)
# fig.tight_layout()
return fig, axdef regions_workflow(fname, sigma=10, cutoff=20):
""" Mostly automated quantification (play with sigma). """
img0 = imread(fname, as_gray=True)
img1 = gaussian(img0, sigma=sigma)
img2 = rescale_intensity(img1, out_range=(-1, 1))
img4 = img1 > threshold_otsu(img1)
label_img = label(1 - img4)
regions = regionprops(label_img)
porosity = 100 * (1 - img4.sum() / img0.size)
contours = find_contours(img4, 0.99)
fig, ax = plot_all_regions(img0, contours, regions, cutoff)
table = regionprops_table(label_img, properties=(
"area",
'axis_major_length',
'axis_minor_length',
'centroid',
"eccentricity",
"equivalent_diameter_area",
"feret_diameter_max",
'orientation',
"perimeter",
"perimeter_crofton"
))
return pd.DataFrame(table)table = regions_workflow(fname)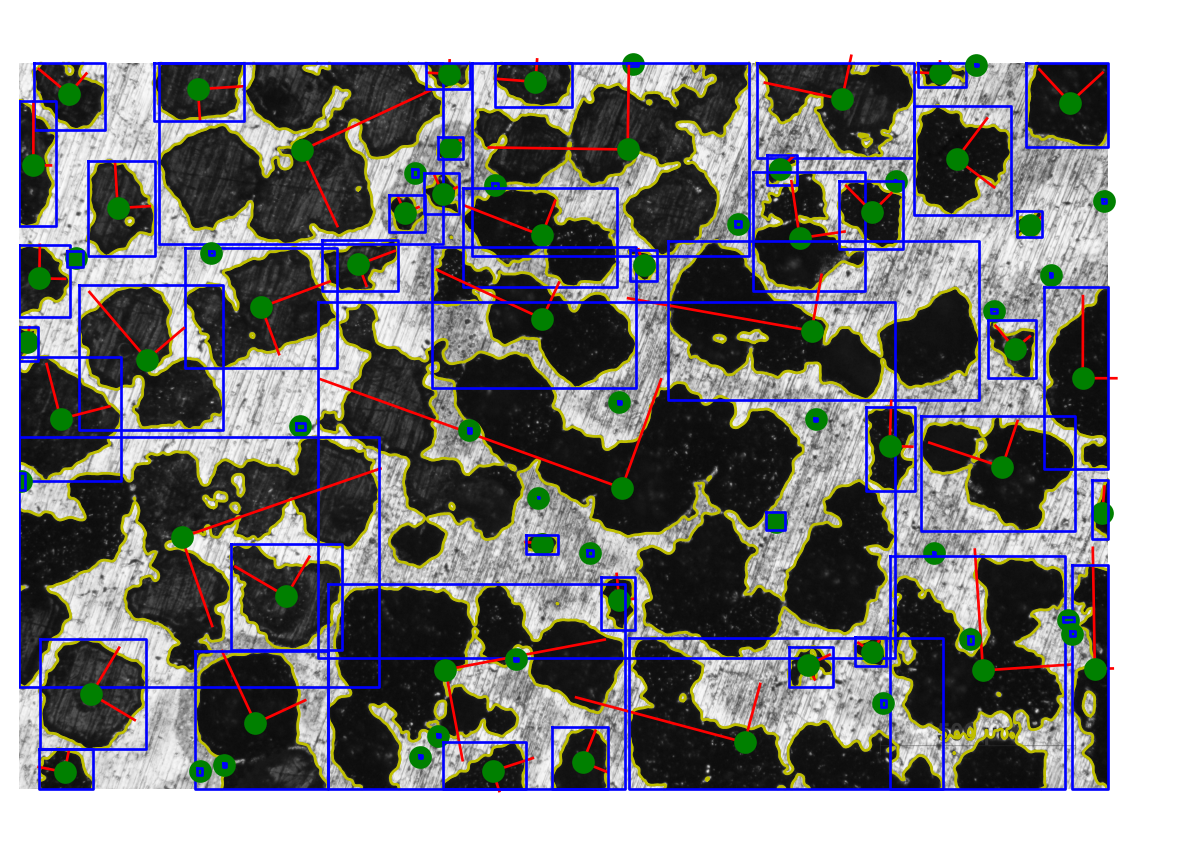
table.head().T| 0 | 1 | 2 | 3 | 4 | |
|---|---|---|---|---|---|
| area | 73161.000000 | 97748.000000 | 665846.000000 | 18612.000000 | 60719.000000 |
| axis_major_length | 376.235279 | 439.661618 | 1353.375215 | 200.600567 | 376.278958 |
| axis_minor_length | 256.600252 | 293.277190 | 796.578811 | 128.380797 | 213.213921 |
| centroid-0 | 148.098618 | 123.927640 | 414.686540 | 50.954760 | 88.785537 |
| centroid-1 | 236.947486 | 850.714746 | 1345.892332 | 2045.305985 | 2456.921507 |
| eccentricity | 0.731333 | 0.745011 | 0.808434 | 0.768390 | 0.823967 |
| equivalent_diameter_area | 305.207271 | 352.784097 | 920.750486 | 153.940035 | 278.046456 |
| feret_diameter_max | 378.498349 | 453.007726 | 1383.046275 | 216.094887 | 368.401954 |
| orientation | 0.875968 | -1.507255 | -1.134392 | 1.518901 | 1.495124 |
| perimeter | 1270.856998 | 1292.482323 | 6860.861720 | 611.297511 | 1013.955411 |
| perimeter_crofton | 1207.529502 | 1228.031596 | 6504.701132 | 582.424248 | 963.971525 |
For post-processing, now you can think about anything, such as creating bins for classification:
table["classes"] = pd.cut(table["area"], bins=4)
table.groupby("classes", observed=True).agg(("count", "mean", "std")).T.head(6)| classes | (-1694.008, 424255.0] | (424255.0, 848507.0] | (1272759.0, 1697011.0] | |
|---|---|---|---|---|
| area | count | 74.000000 | 7.000000 | 1.000000e+00 |
| mean | 54913.635135 | 604669.142857 | 1.697011e+06 | |
| std | 80866.246036 | 137960.255980 | NaN | |
| axis_major_length | count | 74.000000 | 7.000000 | 1.000000e+00 |
| mean | 259.567942 | 1544.173949 | 3.048717e+03 | |
| std | 281.790113 | 293.199681 | NaN |